Julia and Python better together
Introduction
Many data scientists with whom I discuss tell me that they like Julia, but there are some functionalities in Python that they like and would want to keep using.
In this post I want to show that if you are in this situation the answer is: it is fine - you can work with Julia and keep using Python packages you like as a part of your Julia workflow.
The post is tested under Julia 1.8.5 and Status PyCall.jl 1.95.1, Conda.jl 1.8.0, PyPlot.jl 2.11.0, and GLM.jl 1.8.1.
Level 1: Popular Python packages have an interface in Julia
Many (if not majority of) Python packages are written in other languages (like C++) and Python is only a wrapper around them. Most Python users do not care (or even think) about it - they focus on getting their projects delivered.
The same approach can be used in Julia. Specifically, Julia package can be just a wrapper around some Python package. Examples of such popular packages are:
- PyPlot.jl wrapping Matplotlib;
- Pandas.jl wrapping Pandas;
- ScikitLearn.jl wrapping scikit-learn;
- Seaborn.jl wrapping Seaborn;
- SymPy.jl wrapping SymPy.
This is an easy scenario as all you need to do is install a Julia package and you are ready to go and use your favorite Python package.
I will give here one example from Matplotlib documentation. I want to reproduce the Python code given here:
# Python code
fig, ax = plt.subplots(figsize=(5, 2.7))
t = np.arange(0.0, 5.0, 0.01)
s = np.cos(2 * np.pi * t)
ax.plot(t, s, lw=2)
ax.annotate('local max', xy=(2, 1), xytext=(3, 1.5),
arrowprops=dict(facecolor='black', shrink=0.05))
ax.set_ylim(-2, 2)
Now let us do the same in Julia using PyPlot.jl:
# Julia code
using PyPlot
fig, ax = plt.subplots(figsize=(5, 2.7))
t = 0.0:0.01:4.99
s = cos.(2 * π * t)
ax.plot(t, s, lw=2)
ax.annotate("local max", xy=(2, 1), xytext=(3, 1.5),
arrowprops=Dict("facecolor" => "black", "shrink" => 0.05))
ax.set_ylim(-2, 2)
As you can see the codes are almost identical. The only differences are related
to syntax. For example ' is replaced by " and dict by Dict.
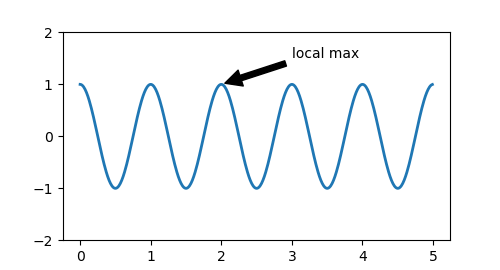
Both codes produce the following plot:
Level 2: You can use any Python package from Julia
Not all Python packages have Julia wrappers. Also, in some cases you might want to work with a different version or configuration of the Python package than provided by the wrapper. Is this a problem? No, this is not a problem at all:
You can load and use any Python package from Julia.
There are two Julia packages that allow for this PyCall.jl and PythonCall.jl. Both packages provide the ability to directly call and fully interoperate with Python from the Julia language. There are some technical differences between them which are described here so that you can decide which one you prefer.
Below I will give you an example of using PyCall.jl.
Assume you like using statsmodels package from Python and would want to reproduce this example from its documentation:
# Python code
import numpy as np
import statsmodels.api as sm
spector_data = sm.datasets.spector.load()
spector_data.exog = sm.add_constant(spector_data.exog, prepend=False)
mod = sm.OLS(spector_data.endog, spector_data.exog)
res = mod.fit()
print(res.summary())
Let me show you how to reproduce it step-by-step in Julia.
We start with loading the packages:
using PyCall
sm = pyimport("statsmodels.api")
Note that using PyCall.jl we can load any Python package using the
pyimport function. The second line might give you an error like this:
julia> sm = pyimport("statsmodels.api")
ERROR: PyError (PyImport_ImportModule
...
This is not a problem. It is just an information that the statsmodels package is not installed under Python. In this case you can easily install it from Julia. Just do:
using Conda
Conda.add("statsmodels")
and you are ready to go. Now sm = pyimport("statsmodels.api") will work.
We are ready to build the model in Julia using statsmodels:
# Julia code
spector_data = sm.datasets.spector.load()
spector_data["exog"] = sm.add_constant(spector_data["exog"], prepend=false)
model = sm.OLS(spector_data["endog"], spector_data["exog"])
res = model.fit()
res.summary()
and you get the output that is the same as in statsmodels documentation:
PyObject <class 'statsmodels.iolib.summary.Summary'>
"""
OLS Regression Results
==============================================================================
Dep. Variable: GRADE R-squared: 0.416
Model: OLS Adj. R-squared: 0.353
Method: Least Squares F-statistic: 6.646
Date: Fri, 17 Feb 2023 Prob (F-statistic): 0.00157
Time: 14:41:35 Log-Likelihood: -12.978
No. Observations: 32 AIC: 33.96
Df Residuals: 28 BIC: 39.82
Df Model: 3
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
GPA 0.4639 0.162 2.864 0.008 0.132 0.796
TUCE 0.0105 0.019 0.539 0.594 -0.029 0.050
PSI 0.3786 0.139 2.720 0.011 0.093 0.664
const -1.4980 0.524 -2.859 0.008 -2.571 -0.425
==============================================================================
Omnibus: 0.176 Durbin-Watson: 2.346
Prob(Omnibus): 0.916 Jarque-Bera (JB): 0.167
Skew: 0.141 Prob(JB): 0.920
Kurtosis: 2.786 Cond. No. 176.
==============================================================================
Notes:
[1] Standard Errors assume that the covariance matrix of the errors
is correctly specified.
Note that the differences in codes are minimal. Again I needed to adjust to
Julia syntax by changing False to false and doing dictionary access using
square brackets like in spector_data["exog"]. All else is identical (and for
this reason I put a comment on top showing which language is used as it could
be easily confused).
You might, however, ask if it is possible to use data from Python in Julia (or data from Julia in Python). Yes - this is also supported. The only thing to remember is that automatic conversion of values between Python and Julia is done for a predefined list of most common types (like arrays, dictionaries).
Let me give an example how this is done by estimating the same regression using GLM.jl. What I will do is transport the data as arrays from Python to Julia (I chose this case as it is most commonly used in my experience).
using GLM
lm(spector_data["exog"].to_numpy(), spector_data["endog"].to_numpy())
And you get the output:
LinearModel{GLM.LmResp{Vector{Float64}}, GLM.DensePredChol{Float64,
LinearAlgebra.CholeskyPivoted{Float64, Matrix{Float64}, Vector{Int64}}}}:
Coefficients:
───────────────────────────────────────────────────────────────────
Coef. Std. Error t Pr(>|t|) Lower 95% Upper 95%
───────────────────────────────────────────────────────────────────
x1 0.463852 0.161956 2.86 0.0078 0.132099 0.795604
x2 0.0104951 0.0194829 0.54 0.5944 -0.0294137 0.0504039
x3 0.378555 0.139173 2.72 0.0111 0.0934724 0.663637
x4 -1.49802 0.523889 -2.86 0.0079 -2.57115 -0.42488
───────────────────────────────────────────────────────────────────
As you can see the results are the same, except that we have lost column names. The reason is that we have used arrays to transport data from Python to Julia (column names also could be transported, but I did not want to complicate the example).
Conclusions
Today the conclusion is short (but for me extremely powerful):
From Julia you have all Julia and all Python packages available to use in your projects.
If you know some package in Python and want to keep using it in Julia it is easy. In most cases you can just copy-paste your Python code to Julia and do minor syntax adjustments and you are done.
I find this interoperability of Julia and Python really amazing.